PairwiseAlignment방법의 하나. 하나의 서열이 다른 하나에 포함되거나 overlap될때의 matches. fragments of genomic DNA sequence들 끼리 비교하거나 더 큰 chromosomal sequence와 비교할 때 필요함. GlobalAlignment의 한 형태이면서 overhanging ends에 벌점을 주지 않는 경우이다.
We want a match to start on the top or left border of the matrix, and finish on the right or bottom border.
Initially
- F(i,0)=0 for i=1,...,n
- F(0,j)=0 for j=1,...,m
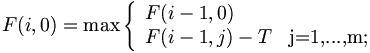

Traceback starts from the maximum point and continues until the top or left edge is reached.
 BioHackersNet
BioHackersNet