PairwiseAlignment방법가운데 하나.
두서열중 하나 혹은 둘다가 길때, 많은 다른 LocalAlignment들이 존재 할 수 있으며, 생물학자역시 이것에 관심이 많다. 예를 들어, many copies of a repeated [Domain] or [Motif]. 이 방법은 asymmetric하며, 하나 혹은 그 이상의 non-overlapping copies of section 들을 찾는다.
multiple matches를 찾는 접근은 Waterman & Eggert 에 의해 1987년 소개됨.
Threshold T. (T=20) --> there are always short LocalAlignment with small positive scores even between entirely unrelated sequences. Usually
- y : containing the domain or motif
- x : the sequence in which we are looking for multiple matches
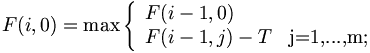
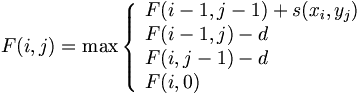
Traceback procedure is g global procedure.
Changing the value of T changes what the algorithm finds
- Increasing T : exclude matches
- Decreasing T : it may split them, as well as finding new weaker ones
 BioHackersNet
BioHackersNet